NGS workflow
Overview
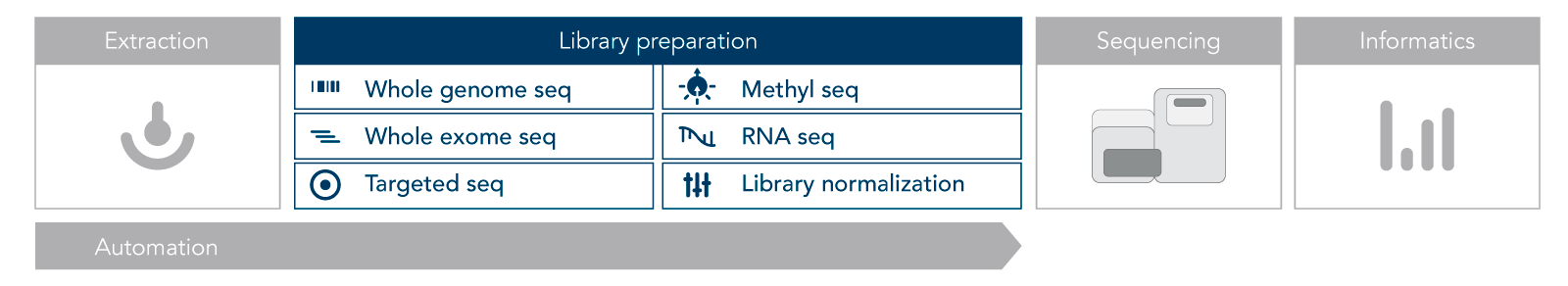
The next generation sequencing workflow consists of four key steps: nucleic acid extraction, library preparation, sequencing, and analysis. The choices you make at each step are critical for your experimental outcomes. Learn more about each step below.
The NGS workflow
NGS Workflow Step 1: Nucleic Acid Isolation
NGS starts with genomic material with either DNA or RNA that is extracted from cells or tissue. Your choice of isolation method or kit is critical to ensure proper lysis of cells and tissues so that you can capture the genetic material in the sample. Selecting the right isolation method or kit will maximize nucleic acid yield, purity, and quality for the next step of NGS library preparation. You should confirm the yield, purity, and quality of your sequences before moving on to the library preparation step.
- Yield: Most library prep methods require nanograms to micrograms of DNA or cDNA (from RNA). Keep this in mind when working with low biomass samples and be sure to select a proper isolation method/kit.
- Purity: Some chemicals contained in nucleic acid isolation kits (phenol, ethanol) and biological materials (heparin, humic acid) can inhibit library preparation. Make sure your nucleic acid isolation method/kit includes steps for removing these inhibitors.
- Quality: The quality of your DNA or RNA should help with a successful library preparation. For example, ensure that the DNA is of high molecular weight and intact, minimize degradation during RNA storage and preparation, and consider using specific isolation methods or kits if you know your starting material is already fragmented.
NGS Workflow Step 2: Library Preparation
The next step in the NGS workflow is library preparation and provides everything you need to do to your isolated, purified genomic material for sequencing. IDT products are compatible with many types of sequencing platforms, including Illumina®, Ion Torrent, MGI, Element Biosciences, Singular Genomics, Ultima Genomics, Pacific Biosciences, and Oxford Nanopore Technologies,
During library prep, nucleic acids are typically fragmented into shorter sequences and adapters are attached to the sequences to make them compatible with the sequencer. Unique identifying sequences called “barcodes” or “indexes” are added to the samples at this stage. Barcodes allow samples to be combined, which is referred to as “pooling” or “multiplexing.” This allows the researcher to perform sequencing simultaneously which saves time and money.
Optional Step: Enrichment
An alternative to whole genome sequencing (WGS) is targeted sequencing, where specific portions of the genome, such as marker genes or other genomic regions of interest, are sequenced in lieu of the entire genome. This approach, called enrichment, is often faster and more cost-effective than sequencing the entire genome. Enrichment can be performed either during library prep (amplicon sequencing) or after library prep (hybridization capture), depending on the enrichment method chosen.
NGS 101 application guide
This detailed overview walks you through major advances in sequencing technology, types of next generation sequencing, their applications and more.
NGS Workflow Step 3: Sequencing
Not sequencing with Illumina? We also help customers design custom solutions for other sequencing platforms. Contact us today to learn more about custom solutions.
Targeted sequencing application guide
This detailed overview walks you through major advances in capture and enrichment technology, types of targeted next generation sequencing, their applications, and more.
NGS Workflow Step 4: Data Analysis
The last step in the NGS workflow is analyzing the sequence reads produced by the sequencer. Just as there are many sequencing methods, there are many data analysis approaches, but they generally fall into one of two categories: read processing and sequence analysis.
- Read processing: read processing is basically a series of quality control steps that clean your sequences and prepare them for downstream analyses. Read processing begins with base calling, where specific nucleotides present at each position in a single sequencing read are identified. Each base call receives a quality score (i.e., how confident are we that the call is correct); reads with low average quality scores can be removed from the dataset. Adaptor sequences added during library prep are then removed, and if barcodes were added, reads can be separated and grouped based on the sample they came from in a process called demultiplexing.
- Sequence analysis: Sequence analysis typically begins by aligning your filtered set of sequences against a reference genome or database. Then, any number of steps can be performed, including removal of duplicate reads, alignment, and assembly (when performing de novo sequencing), genome annotation, variant calling/annotation, determination of gene counts, and more. There are a wide variety of bioinformatics tools and data analysis apps you can use to analyze your sequences, each with its own benefits and caveats.
Get started with next generation sequencing
Whether you need a library prep kit, custom NGS adaptors, or SARS-CoV2 research primers and probes, we have it. Talk to a scientific application specialist today to design and customize your NGS workflow.
